python操作txt檔案中資料教程[2]-python提取txt檔案
阿新 • • 發佈:2018-11-26
python操作txt檔案中資料教程[2]-python提取txt檔案中的行列元素
覺得有用的話,歡迎一起討論相互學習~Follow Me
- 原始txt檔案
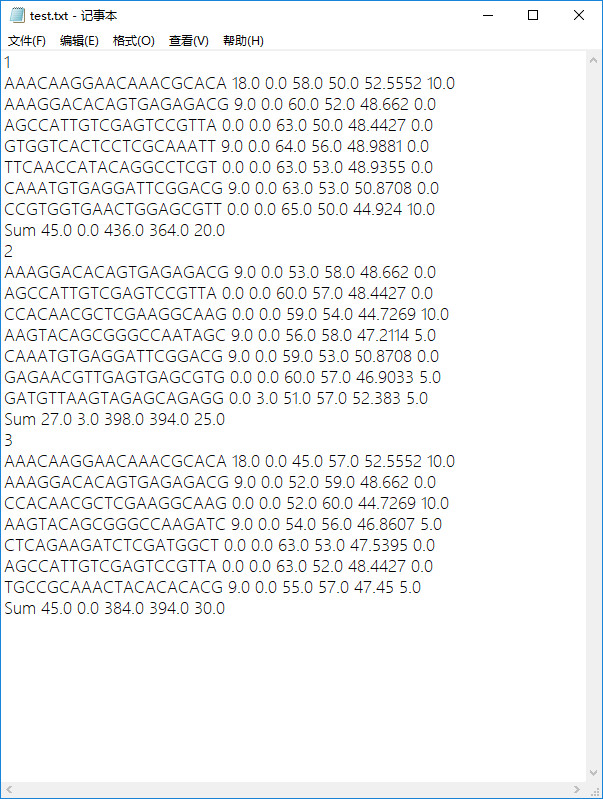
程式實現後結果-將txt中元素提取並儲存在csv中
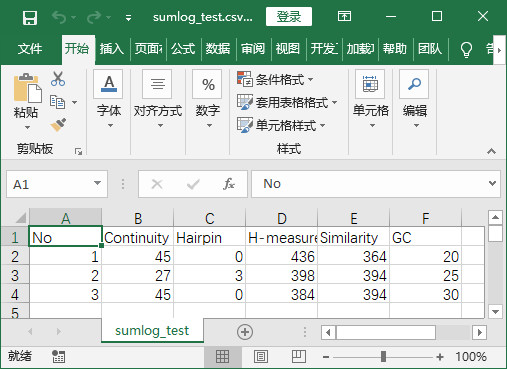
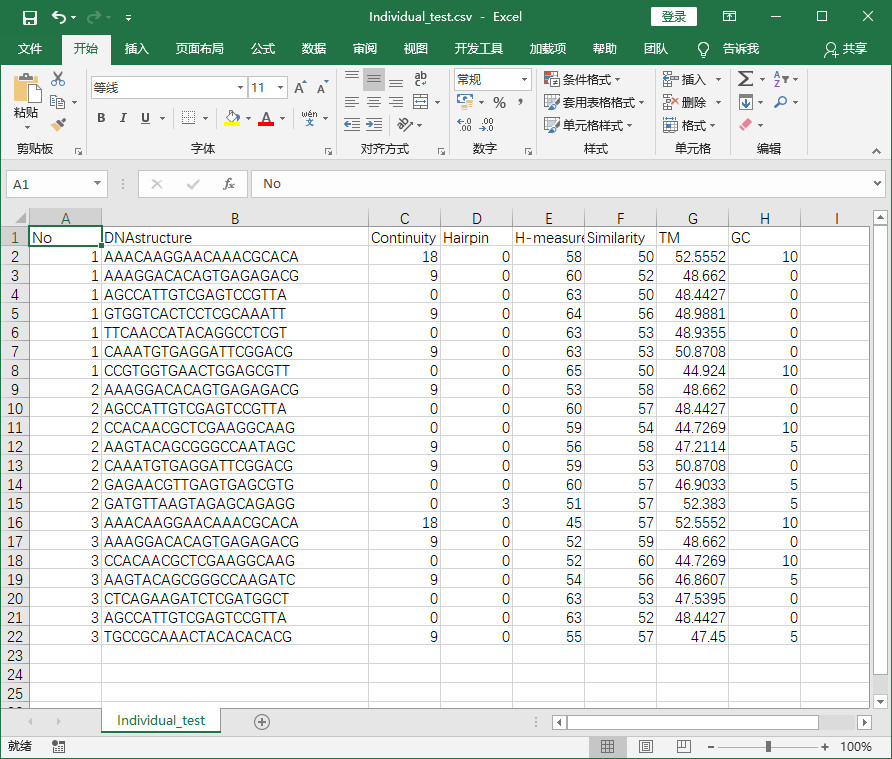
程式實現
import csv filename = "./test/test.txt" Sum_log_file = "./test/sumlog_test.csv" Individual_log_file = "./test/Individual_test.csv" DNA_log = [] # 精英種群個體日誌mod9=1-8 Sum_log = [] # 精英種群總體日誌mod9=0 DNA_Group = 7 # 表示每7條DNA組成一個組 # NO+'Sum 45.0 0.0 436.0 364.0 20.0\n'中屬性一共6個屬性,,則設為8列的二維陣列 sum_evaindex = [[] for i in range(6)] # 個體有8個屬性,則設為8列的二維陣列 Individual_evaindex = [[] for i in range(8)] # 將txt中檔案資訊儲存到Sum_log和DNA_log列表中 with open(filename, 'r') as f: i = 1 for line in f.readlines(): if i%9 == 0: Sum_log.append(line) else: DNA_log.append(line) i = i + 1 f.close() # print(Sum_log) # print(DNA_log) # ['Sum 45.0 0.0 436.0 364.0 20.0\n', 'Sum 27.0 3.0 398.0 394.0 25.0\n', 'Sum 45.0 0.0 384.0 394.0 30.0'] # ['1\n', 'AAACAAGGAACAAACGCACA 18.0 0.0 58.0 50.0 52.5552 10.0\n', 'AAAGGACACAGTGAGAGACG 9.0 0.0 60.0 52.0 48.662 0.0\n', # 'AGCCATTGTCGAGTCCGTTA 0.0 0.0 63.0 50.0 48.4427 0.0\n', 'GTGGTCACTCCTCGCAAATT 9.0 0.0 64.0 56.0 48.9881 0.0\n', # 'TTCAACCATACAGGCCTCGT 0.0 0.0 63.0 53.0 48.9355 0.0\n', 'CAAATGTGAGGATTCGGACG 9.0 0.0 63.0 53.0 50.8708 0.0\n', # 'CCGTGGTGAACTGGAGCGTT 0.0 0.0 65.0 50.0 44.924 10.0\n', '2\n', 'AAAGGACACAGTGAGAGACG 9.0 0.0 53.0 58.0 48.662 0.0\n', # 'AGCCATTGTCGAGTCCGTTA 0.0 0.0 60.0 57.0 48.4427 0.0\n', 'CCACAACGCTCGAAGGCAAG 0.0 0.0 59.0 54.0 44.7269 10.0\n', # 'AAGTACAGCGGGCCAATAGC 9.0 0.0 56.0 58.0 47.2114 5.0\n', 'CAAATGTGAGGATTCGGACG 9.0 0.0 59.0 53.0 50.8708 0.0\n', # 'GAGAACGTTGAGTGAGCGTG 0.0 0.0 60.0 57.0 46.9033 5.0\n', 'GATGTTAAGTAGAGCAGAGG 0.0 3.0 51.0 57.0 52.383 5.0\n', '3\n', # 'AAACAAGGAACAAACGCACA 18.0 0.0 45.0 57.0 52.5552 10.0\n', 'AAAGGACACAGTGAGAGACG 9.0 0.0 52.0 59.0 48.662 0.0\n', # 'CCACAACGCTCGAAGGCAAG 0.0 0.0 52.0 60.0 44.7269 10.0\n', 'AAGTACAGCGGGCCAAGATC 9.0 0.0 54.0 56.0 46.8607 5.0\n', # 'CTCAGAAGATCTCGATGGCT 0.0 0.0 63.0 53.0 47.5395 0.0\n', 'AGCCATTGTCGAGTCCGTTA 0.0 0.0 63.0 52.0 48.4427 0.0\n', # 'TGCCGCAAACTACACACACG 9.0 0.0 55.0 57.0 47.45 5.0\n'] # 遍歷行,並將列屬性儲存到對應列中 Sum_no = 1 for Sum in Sum_log: # print(Sum.split("\n")[0].split(" ")[1:]) # ['45.0', '0.0', '436.0', '364.0', '20.0'] # ['27.0', '3.0', '398.0', '394.0', '25.0'] # ['45.0', '0.0', '384.0', '394.0', '30.0'] sum_eva_index = Sum.split("\n")[0].split(" ")[1:] sum_evaindex[0].append(int(Sum_no)) sum_evaindex[1].append(float(sum_eva_index[0])) # Con sum_evaindex[2].append(float(sum_eva_index[1])) # HP sum_evaindex[3].append(float(sum_eva_index[2])) # Hm sum_evaindex[4].append(float(sum_eva_index[3])) # Si sum_evaindex[5].append(float(sum_eva_index[4])) # GC Sum_no = Sum_no + 1 # print(sum_evaindex[0]) # [45.0, 27.0, 45.0] # 遍歷個體資訊,並將其儲存到Individual_evaindex列表中 dna_log_no = 0 for dna_log in DNA_log: if (dna_log_no + 1)%8 == 1: # print(int(dna_log.split("\n")[0])) # 以列儲存序號值,並且重複DNA_Group次 for i in range(DNA_Group): Individual_evaindex[0].append(int(dna_log.split("\n")[0])) else: Individual_evaindex[1].append(dna_log.split("\n")[0].split(" ")[0]) # 所有DNA序列全部記載,使用原有的str字串型別記載 Individual_evaindex[2].append(float(dna_log.split("\n")[0].split(" ")[1])) # DNA序列的連續值Con,注意要轉換為浮點數型別 Individual_evaindex[3].append(float(dna_log.split("\n")[0].split(" ")[2])) # Hp莖區匹配 Individual_evaindex[4].append(float(dna_log.split("\n")[0].split(" ")[3])) # H-measure Individual_evaindex[5].append(float(dna_log.split("\n")[0].split(" ")[4])) # Similarity Individual_evaindex[6].append(float(dna_log.split("\n")[0].split(" ")[5])) # TM Individual_evaindex[7].append(float(dna_log.split("\n")[0].split(" ")[6])) # GC dna_log_no = dna_log_no + 1 # print(Individual_evaindex[0]) #[1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3] # print(Individual_evaindex[1]) # print(Individual_evaindex[2]) # print(Individual_evaindex[3]) # print(Individual_evaindex[4]) # print(Individual_evaindex[5]) # print(Individual_evaindex[6]) # print(Individual_evaindex[7]) # ['AAACAAGGAACAAACGCACA', 'AAAGGACACAGTGAGAGACG', 'AGCCATTGTCGAGTCCGTTA', 'GTGGTCACTCCTCGCAAATT', 'TTCAACCATACAGGCCTCGT', # 'CAAATGTGAGGATTCGGACG', 'CCGTGGTGAACTGGAGCGTT', 'AAAGGACACAGTGAGAGACG', 'AGCCATTGTCGAGTCCGTTA', 'CCACAACGCTCGAAGGCAAG', # 'AAGTACAGCGGGCCAATAGC', 'CAAATGTGAGGATTCGGACG', 'GAGAACGTTGAGTGAGCGTG', 'GATGTTAAGTAGAGCAGAGG', 'AAACAAGGAACAAACGCACA', # 'AAAGGACACAGTGAGAGACG', 'CCACAACGCTCGAAGGCAAG', 'AAGTACAGCGGGCCAAGATC', 'CTCAGAAGATCTCGATGGCT', 'AGCCATTGTCGAGTCCGTTA', # 'TGCCGCAAACTACACACACG'] # [18.0, 9.0, 0.0, 9.0, 0.0, 9.0, 0.0, 9.0, 0.0, 0.0, 9.0, 9.0, 0.0, 0.0, 18.0, 9.0, 0.0, 9.0, 0.0, 0.0, 9.0] # [0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 3.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0] # [58.0, 60.0, 63.0, 64.0, 63.0, 63.0, 65.0, 53.0, 60.0, 59.0, 56.0, 59.0, 60.0, 51.0, 45.0, 52.0, 52.0, 54.0, 63.0, 63.0, # 55.0] # [50.0, 52.0, 50.0, 56.0, 53.0, 53.0, 50.0, 58.0, 57.0, 54.0, 58.0, 53.0, 57.0, 57.0, 57.0, 59.0, 60.0, 56.0, 53.0, 52.0, # 57.0] # [52.5552, 48.662, 48.4427, 48.9881, 48.9355, 50.8708, 44.924, 48.662, 48.4427, 44.7269, 47.2114, 50.8708, 46.9033, # 52.383, 52.5552, 48.662, 44.7269, 46.8607, 47.5395, 48.4427, 47.45] # [10.0, 0.0, 0.0, 0.0, 0.0, 0.0, 10.0, 0.0, 0.0, 10.0, 5.0, 0.0, 5.0, 5.0, 10.0, 0.0, 10.0, 5.0, 0.0, 0.0, 5.0] Sum_log_file_header = ["No", "Continuity", "Hairpin", "H-measure", "Similarity", "GC"] # 將資料寫入csv日誌檔案中 with open(Sum_log_file, "w", newline='') as f: writer = csv.writer(f) writer.writerow(Sum_log_file_header) # 注意,此處使用writerow而不是使用writerows for i in range(sum_evaindex[0][-1]): # i 取(0,1,2) writer.writerow( [sum_evaindex[0][i], sum_evaindex[1][i], sum_evaindex[2][i], sum_evaindex[3][i], sum_evaindex[4][i], sum_evaindex[5][i]]) f.close() Individual_log_file_header = ["No", "DNAstructure", "Continuity", "Hairpin", "H-measure", "Similarity", "TM", "GC"] with open(Individual_log_file, "w", newline='') as f: writer = csv.writer(f) writer.writerow(Individual_log_file_header) # 注意,此處使用writerow而不是使用writerows for i in range(sum_evaindex[0][-1]*DNA_Group): writer.writerow( [Individual_evaindex[0][i], Individual_evaindex[1][i], Individual_evaindex[2][i], Individual_evaindex[3][i], Individual_evaindex[4][i], Individual_evaindex[5][i], Individual_evaindex[6][i], Individual_evaindex[7][i]]) f.close()
測試版本
filename = "./test.txt" DNA_log = [] # 精英種群個體日誌mod9=2-8 Sum_log = [] # 精英種群總體日誌mod9=0 Num_log = [] # 序號日誌mod9=1 Num_int = [] # 擷取序號為int型別 sum_evaindex = [[] for i in range(5)] Individual_evaindex = [[] for i in range(8)] with open(filename, 'r') as f: i = 1 for line in f.readlines(): if i%9 == 1: Num_log.append(line) elif i%9 == 0: Sum_log.append(line) else: DNA_log.append(line) i = i + 1 f.close() print(Num_log) print(Num_log[1]) # 其中存著的不是數字1,而是字串'2\n',所以會有空行的情況 # ['1\n', '2\n', '3\n'] # 2 # # print(Sum_log) print(DNA_log) # ['Sum 45.0 0.0 436.0 364.0 20.0\n', 'Sum 27.0 3.0 398.0 394.0 25.0\n', 'Sum 45.0 0.0 384.0 394.0 30.0'] # ['AAACAAGGAACAAACGCACA 18.0 0.0 58.0 50.0 52.5552 10.0\n', 'AAAGGACACAGTGAGAGACG 9.0 0.0 60.0 52.0 48.662 0.0\n', # 'AGCCATTGTCGAGTCCGTTA 0.0 0.0 63.0 50.0 48.4427 0.0\n', 'GTGGTCACTCCTCGCAAATT 9.0 0.0 64.0 56.0 48.9881 0.0\n', # 'TTCAACCATACAGGCCTCGT 0.0 0.0 63.0 53.0 48.9355 0.0\n', 'CAAATGTGAGGATTCGGACG 9.0 0.0 63.0 53.0 50.8708 0.0\n', # 'CCGTGGTGAACTGGAGCGTT 0.0 0.0 65.0 50.0 44.924 10.0\n', 'AAAGGACACAGTGAGAGACG 9.0 0.0 53.0 58.0 48.662 0.0\n', # 'AGCCATTGTCGAGTCCGTTA 0.0 0.0 60.0 57.0 48.4427 0.0\n', 'CCACAACGCTCGAAGGCAAG 0.0 0.0 59.0 54.0 44.7269 10.0\n', # 'AAGTACAGCGGGCCAATAGC 9.0 0.0 56.0 58.0 47.2114 5.0\n', 'CAAATGTGAGGATTCGGACG 9.0 0.0 59.0 53.0 50.8708 0.0\n', # 'GAGAACGTTGAGTGAGCGTG 0.0 0.0 60.0 57.0 46.9033 5.0\n', 'GATGTTAAGTAGAGCAGAGG 0.0 3.0 51.0 57.0 52.383 5.0\n', # 'AAACAAGGAACAAACGCACA 18.0 0.0 45.0 57.0 52.5552 10.0\n', 'AAAGGACACAGTGAGAGACG 9.0 0.0 52.0 59.0 48.662 0.0\n', # 'CCACAACGCTCGAAGGCAAG 0.0 0.0 52.0 60.0 44.7269 10.0\n', 'AAGTACAGCGGGCCAAGATC 9.0 0.0 54.0 56.0 46.8607 5.0\n', # 'CTCAGAAGATCTCGATGGCT 0.0 0.0 63.0 53.0 47.5395 0.0\n', 'AGCCATTGTCGAGTCCGTTA 0.0 0.0 63.0 52.0 48.4427 0.0\n', # 'TGCCGCAAACTACACACACG 9.0 0.0 55.0 57.0 47.45 5.0\n'] for no in Num_log: # print(no[0]) # 字元形式的數字1,這是錯的,因為有可能序號超過一位數 # Num_int.append(int(no.split("\n"))) ['1', ''] Num_int.append(int(no.split("\n")[0])) for Sum in Sum_log: # print(Sum.split("\n")[0].split(" ")[1:]) # ['45.0', '0.0', '436.0', '364.0', '20.0'] # ['27.0', '3.0', '398.0', '394.0', '25.0'] # ['45.0', '0.0', '384.0', '394.0', '30.0'] sum_eva_index = Sum.split("\n")[0].split(" ")[1:] sum_evaindex[0].append(float(sum_eva_index[0])) sum_evaindex[1].append(float(sum_eva_index[1])) sum_evaindex[2].append(float(sum_eva_index[2])) sum_evaindex[3].append(float(sum_eva_index[3])) sum_evaindex[4].append(float(sum_eva_index[4])) print(sum_evaindex[0]) # [45.0, 27.0, 45.0]
