第3章 特徵選擇與特徵工程
阿新 • • 發佈:2019-01-05
標籤編碼,字典向量化,特徵雜湊
LabelEncoder和OneHotEncoder 在特徵工程中的應用
對於性別,sex,一般的屬性值是male和female。兩個值。那麼不靠譜的方法直接用0表示male,用1表示female 了。所以要用one-hot編碼。
array([[0., 1.],
[1., 0.],
[0., 1.],
[0., 1.],
[1., 0.],
[1., 0.]])
classes_:
Holds the label for each class
>>> from from Label encoding
[0 0 0 1 0 1 1 0 0 1]
Label decoding
['Male', 'Female', 'Male', 'Male', 'Female', 'Female']
Label binarization
[[0]
[0]
[0]
[1]
[0]
[1]
[1]
[0]
[0]
[1]]
Label decoding
Dictionary data vectorization
[[10. 15. 0. 0.]
[-5. 0. 22. 0.]
[ 0. 0. -2. 10.]]
Vocabulary:
{'feature_1': 0, 'feature_2': 1, 'feature_3': 2, 'feature_4': 3}
Feature hashing
Feature decoding
[[0. 0. 0. ... 0. 0. 0.]
[0. 0. 0. ... 0. 0. 0.]
[0. 0. 0. ... 0. 0. 0.]]
[[1., 0., 0., 1., 0., 0., 0., 1., 0.]
[0., 1., 0., 0., 1., 0., 0., 0., 1.]
[0., 1., 1., 0., 0., 0., 0., 1., 0.]
[1., 0., 0., 0., 0., 1., 0., 1., 0.]
[1., 0., 0., 0., 0., 0., 1., 0., 1.]]
處理缺失資料
from __future__ import print_function
import numpy as np
from sklearn.preprocessing import Imputer
# For reproducibility
np.random.seed(1000)
if __name__ == '__main__':
data = np.array([[1, np.nan, 2], [2, 3, np.nan], [-1, 4, 2]])
print(data)
# Imputer with mean-strategy
print('Mean strategy')
imp = Imputer(strategy='mean')
print(imp.fit_transform(data))
# Imputer with median-strategy
print('Median strategy')
imp = Imputer(strategy='median')
print(imp.fit_transform(data))
# Imputer with most-frequent-strategy
print('Most-frequent strategy')
imp = Imputer(strategy='most_frequent')
print(imp.fit_transform(data))
[[ 1. nan 2.]
[ 2. 3. nan]
[-1. 4. 2.]]
Mean strategy
[[ 1. 3.5 2. ]
[ 2. 3. 2. ]
[-1. 4. 2. ]]
Median strategy
[[ 1. 3.5 2. ]
[ 2. 3. 2. ]
[-1. 4. 2. ]]
Most-frequent strategy
[[ 1. 3. 2.]
[ 2. 3. 2.]
[-1. 4. 2.]]
資料標準化
標準化(Standardization):對資料的分佈的進行轉換,使其符合某種分佈(比如正態分佈)的一種非線性特徵變換。
method
2.1 Rescaling (min-max normalization)
2.2 Mean normalization
2.3 Standardization
2.4 Scaling to unit length
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler, RobustScaler
# For reproducibility
np.random.seed(1000)
if __name__ == '__main__':
# Create a dummy dataset
data = np.ndarray(shape=(100, 2))
for i in range(100):
data[i, 0] = 2.0 + np.random.normal(1.5, 3.0)
data[i, 1] = 0.5 + np.random.normal(1.5, 3.0)
# Show the original and the scaled dataset
fig, ax = plt.subplots(1, 2, figsize=(14, 5))
ax[0].scatter(data[:, 0], data[:, 1])
ax[0].set_xlim([-10, 10])
ax[0].set_ylim([-10, 10])
ax[0].grid()
ax[0].set_xlabel('X')
ax[0].set_ylabel('Y')
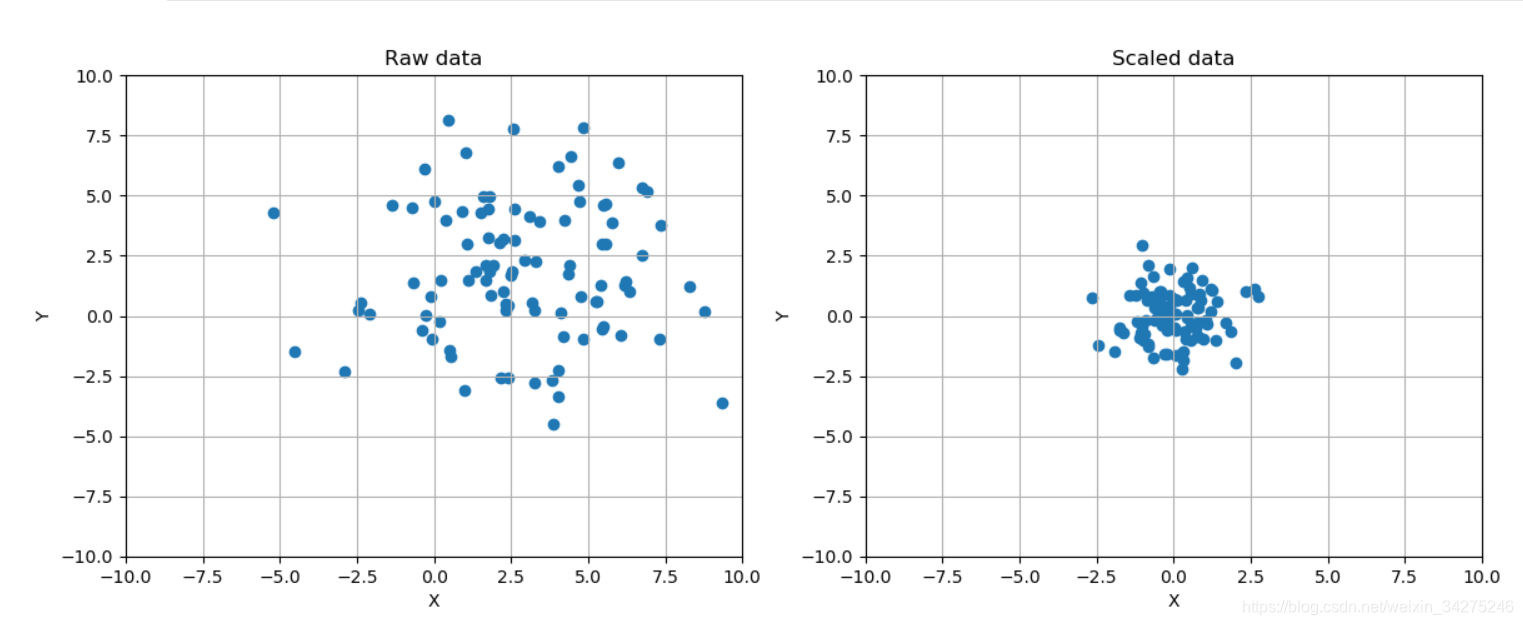
ax[0].set_title('Raw data')
# Scale data
ss = StandardScaler()
scaled_data = ss.fit_transform(data)
ax[1].scatter(scaled_data[:, 0], scaled_data[:, 1])
ax[1].set_xlim([-10, 10])
ax[1].set_ylim([-10, 10])
ax[1].grid()
ax[1].set_xlabel('X')
ax[1].set_ylabel('Y')
ax[1].set_title('Scaled data')
plt.show()
# Scale data using a Robust Scaler
fig, ax = plt.subplots(2, 2, figsize=(8, 8))
ax[0, 0].scatter(data[:, 0], data[:, 1])
ax[0, 0].set_xlim([-10, 10])
ax[0, 0].set_ylim([-10, 10])
ax[0, 0].grid()
ax[0, 0].set_xlabel('X')
ax[0, 0].set_ylabel('Y')
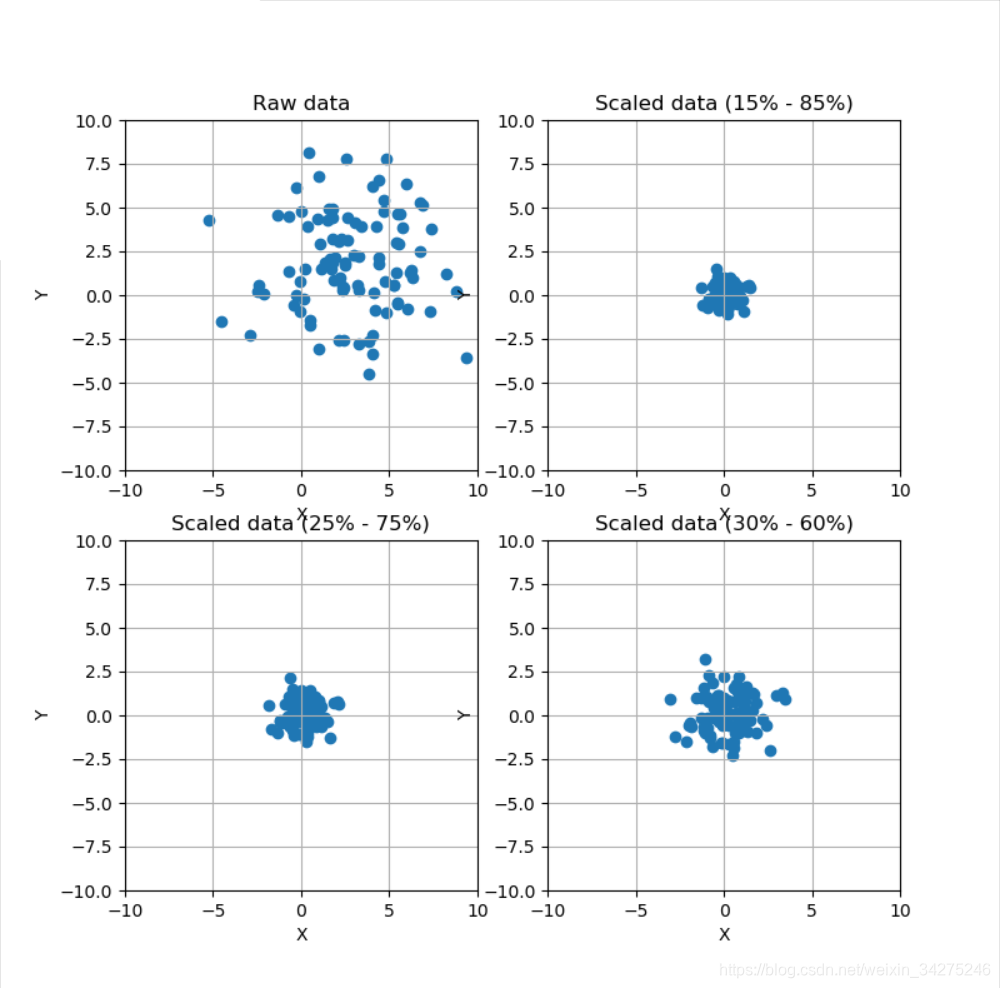
ax[0, 0].set_title('Raw data')
rs = RobustScaler(quantile_range=(15, 85))
scaled_data = rs.fit_transform(data)
ax[0, 1].scatter(scaled_data[:, 0], scaled_data[:, 1])
ax[0, 1].set_xlim([-10, 10])
ax[0, 1].set_ylim([-10, 10])
ax[0, 1].grid()
ax[0, 1].set_xlabel('X')
ax[0, 1].set_ylabel('Y')
ax[0, 1].set_title('Scaled data (15% - 85%)')
rs1 = RobustScaler(quantile_range=(25, 75))
scaled_data1 = rs1.fit_transform(data)
ax[1, 0].scatter(scaled_data1[:, 0], scaled_data1[:, 1])
ax[1, 0].set_xlim([-10, 10])
ax[1, 0].set_ylim([-10, 10])
ax[1, 0].grid()
ax[1, 0].set_xlabel('X')
ax[1, 0].set_ylabel('Y')
ax[1, 0].set_title('Scaled data (25% - 75%)')
rs2 = RobustScaler(quantile_range=(30, 65))
scaled_data2 = rs2.fit_transform(data)
ax[1, 1].scatter(scaled_data2[:, 0], scaled_data2[:, 1])
ax[1, 1].set_xlim([-10, 10])
ax[1, 1].set_ylim([-10, 10])
ax[1, 1].grid()
ax[1, 1].set_xlabel('X')
ax[1, 1].set_ylabel('Y')
ax[1, 1].set_title('Scaled data (30% - 60%)')
plt.show()
Compare the effect of different scalers on data with outliers
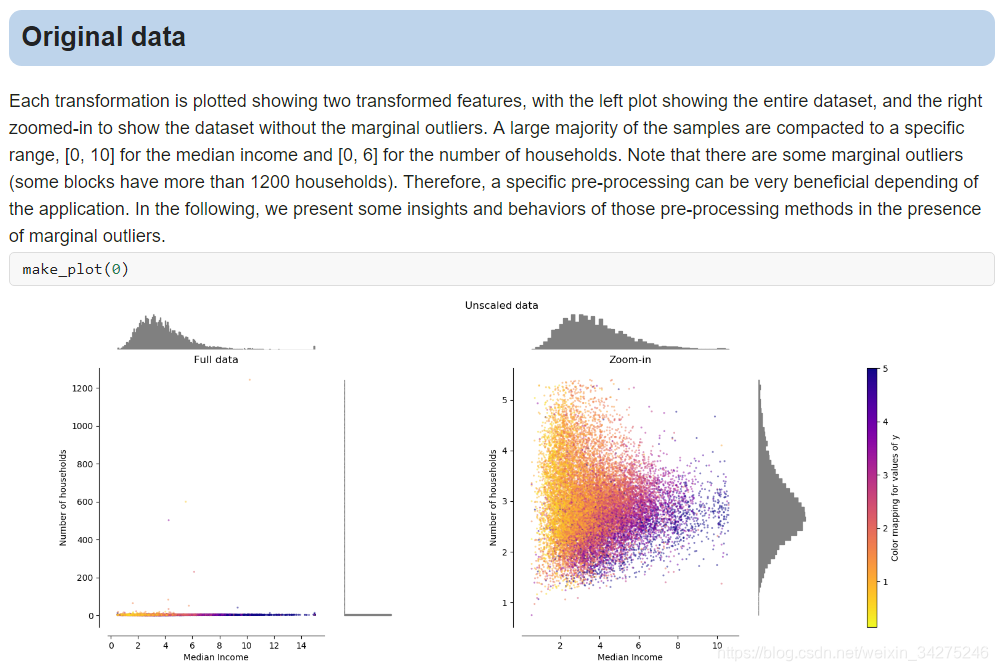
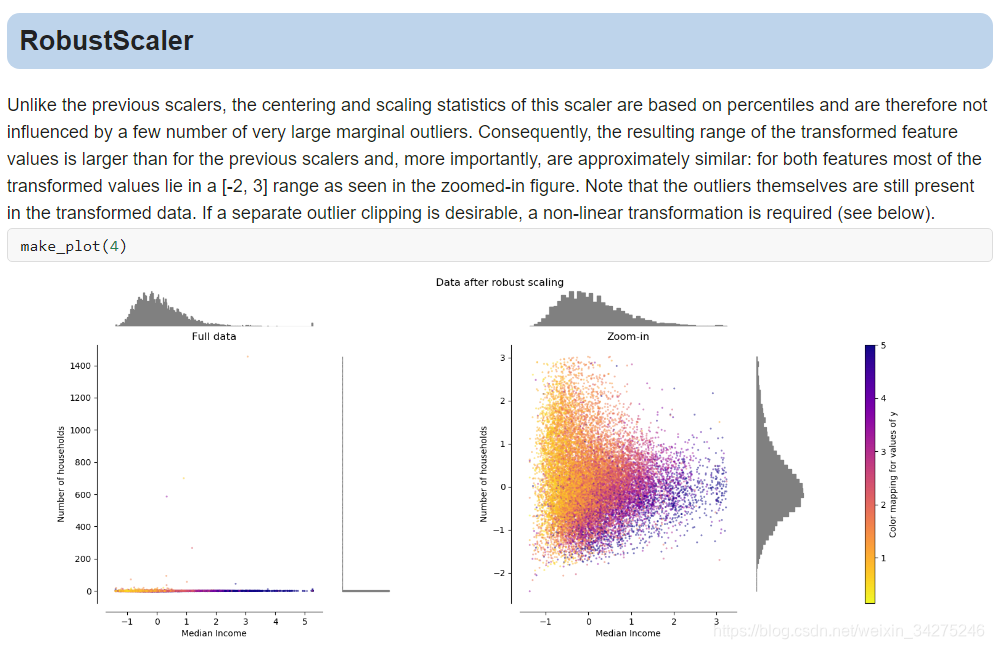
資料歸一化
歸一化:對資料的數值範圍進行特定縮放,但不改變其資料分佈的一種線性特徵變換。
1.min-max 歸一化:將數值範圍縮放到(0,1),但沒有改變資料分佈;
2.z-score 歸一化:將數值範圍縮放到0附近, 但沒有改變資料分佈;
from __future__ import print_function
import numpy as np
from sklearn.preprocessing import Normalizer
# For reproducibility
np.random.seed(1000)
if __name__ == '__main__':
# Create a dummy dataset
data = np.array([1.0, 2.0, 8])
print(data)
# Max normalization
n_max = Normalizer(norm='max')
nm = n_max.fit_transform(data.reshape(1, -1))
print(nm)
# L1 normalization
n_l1 = Normalizer(norm='l1')
nl1 = n_l1.fit_transform(data.reshape(1, -1))
print(nl1)
# L2 normalization
n_l2 = Normalizer(norm='l2')
nl2 = n_l2.fit_transform(data.reshape(1, -1))
print(nl2)
[1. 2. 8.]
[[0.125 0.25 1. ]]
[[0.09090909 0.18181818 0.72727273]]
[[0.12038585 0.24077171 0.96308682]]
特徵篩選
from __future__ import print_function
import numpy as np
from sklearn.datasets import load_boston, load_iris
from sklearn.feature_selection import SelectKBest, SelectPercentile, chi2, f_regression
# For reproducibility
np.random.seed(1000)
if __name__ == '__main__':
# Load Boston data
regr_data = load_boston()
print('Boston data shape')
print(regr_data.data.shape)
# Select the best k features with regression test
kb_regr = SelectKBest(f_regression)
X_b = kb_regr.fit_transform(regr_data.data, regr_data.target)
print('K-Best-filtered Boston dataset shape')
print(X_b.shape)
print('K-Best scores')
print(kb_regr.scores_)
# Load iris data
class_data = load_iris()
print('Iris dataset shape')
print(class_data.data.shape)
# Select the best k features using Chi^2 classification test
perc_class = SelectPercentile(chi2, percentile=15)
X_p = perc_class.fit_transform(class_data.data, class_data.target)
print('Chi2-filtered Iris dataset shape')
print(X_p.shape)
print('Chi2 scores')
print(perc_class.scores_)
Boston data shape
(506, 13)
K-Best-filtered Boston dataset shape
(506, 10)
K-Best scores
[ 88.15124178 75.2576423 153.95488314 15.97151242 112.59148028
471.84673988 83.47745922 33.57957033 85.91427767 141.76135658
175.10554288 63.05422911 601.61787111]
Iris dataset shape
(150, 4)
Chi2-filtered Iris dataset shape
(150, 1)
Chi2 scores
[ 10.81782088 3.59449902 116.16984746 67.24482759]
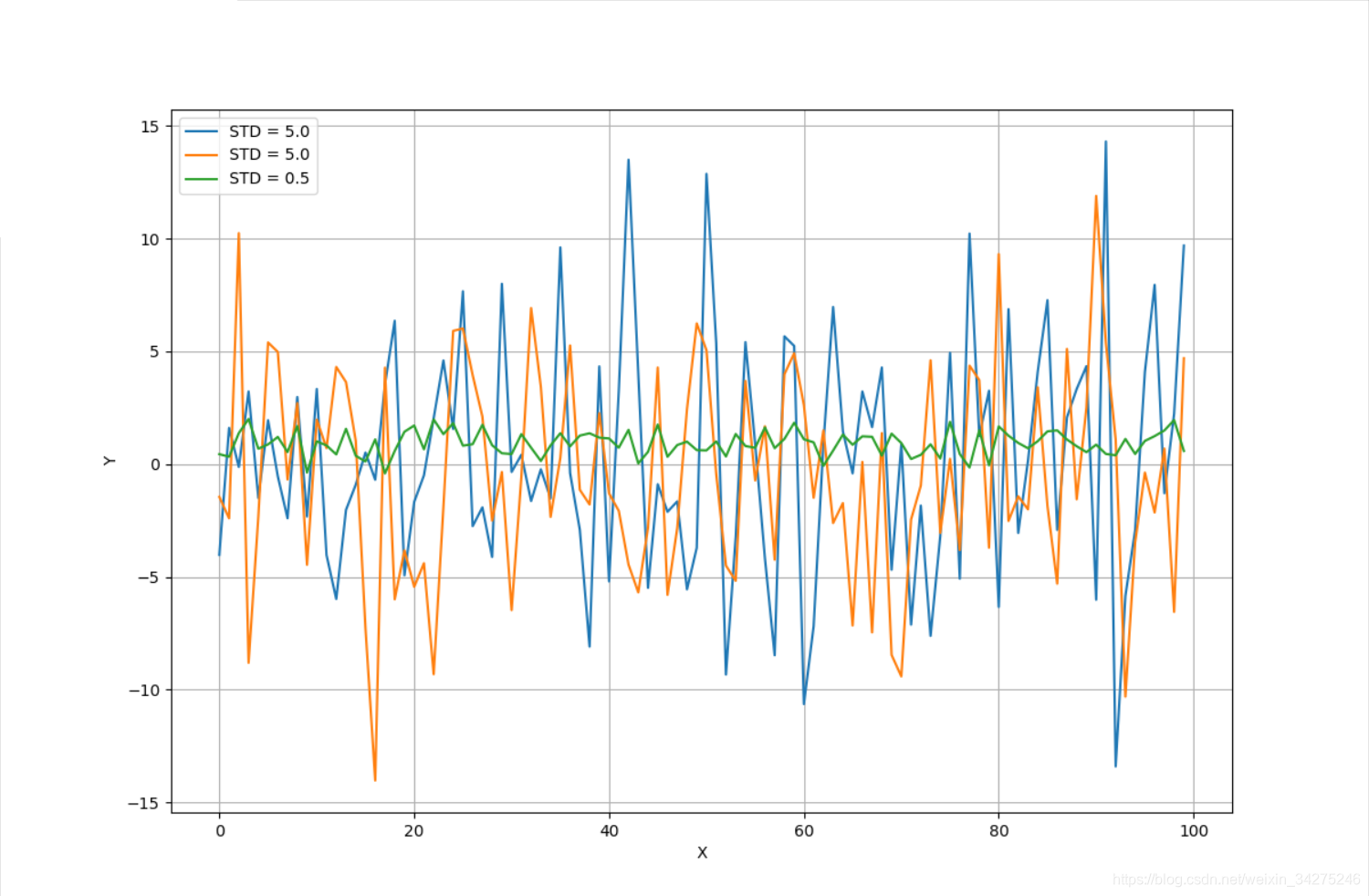
特徵選擇
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
from sklearn.feature_selection import VarianceThreshold
# For reproducibility
np.random.seed(1000)
if __name__ == '__main__':
# Create a dummy dataset
X = np.ndarray(shape=(100, 3))
X[:, 0] = np.random.normal(0.0, 5.0, size=100)
X[:, 1] = np.random.normal(0.5, 5.0, size=100)
X[:, 2] = np.random.normal(1.0, 0.5, size=100)
# Show the dataset
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
ax.grid()
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.plot(X[:, 0], label='STD = 5.0')
ax.plot(X[:, 1], label='STD = 5.0')
ax.plot(X[:, 2], label='STD = 0.5')
plt.legend()
plt.show()
# Impose a variance threshold
print('Samples before variance thresholding')
print(X[0:3, :])
vt = VarianceThreshold(threshold=1.5)
X_t = vt.fit_transform(X)
# After the filter has removed the componenents
print('Samples after variance thresholding')
print(X_t[0:3, :])