蛋白質硝基化修飾位點預測工具 —— GPS-YNO2 1.0
|
Zexian Liu , Jun Cao, Qian Ma, Xinjiao Gao, Jian Ren and Yu Xue . (2010) GPS-YNO2: Computational prediction of tyrosine nitration sites in proteins. Molecular BioSystems (In press). |
![]()
Early after the discovery of the freely-diffusible signaling molecule and second messenger nitric oxide
(NO)
In this work, we manually collected 1066 experimentally indentified protein nitration sites in 554 unique proteins from scientific literature. A previously self-developed GPS (Group-based Prediction System) algorithm was employed with great improvement. We calculated the leave-one-out validation and 4-, 6-, 8-, 10-fold cross-validations to evaluate the prediction performance and system robustness. The leave-one-out validation result is accuracy (Ac ) of 76.51% , sensitivity (Sn ) of 50.09% , and specificity (Sp ) of 80.18% . T he online service and stand-alone packages of GPS-YNO2 1.0 were implemented in JAVA 1.4.2 and freely available at: http://yno2.biocuckoo.org/ .
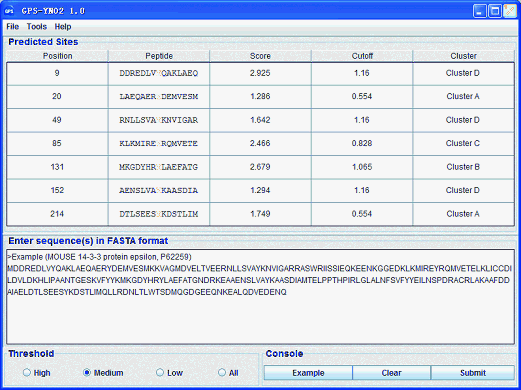
GPS-YNO2 1.0 User Interface
